Eukaryote and prokaryote chromosomes and gene expression
Eukaryotic chromosome organization
Chromosomal proteins
- Histones: Form octamers that DNA wraps around, creating the repeating “beads on a string” pattern. This wrapping is a major contributor to chromatin compaction.
- Nonhistone chromosomal proteins: Include all other chromosome-associated proteins. They serve many functions, including regulation, enzymatic activity, and structural support.
Single-copy versus repetitive DNA
- Some DNA sequences appear only once (single-copy), while others occur in many copies (repetitive DNA).
- Tandem repeats are repeated sequences placed back-to-back (e.g., CAGCAGCAG).
- Excessive trinucleotide repeats can cause disorders such as Huntington disease or fragile X syndrome.
- Satellites, minisatellites, microsatellites: Clusters of repeated sequences. Their characteristic patterns can be used for paternity testing, forensic analysis, or confirming bone marrow engraftment by matching donor and recipient DNA.
- Interspersed repeats: Produced by mobile genetic elements (such as transposons). These elements often contain repetitive DNA at both ends and can move to new locations within the genome.
Supercoiling
Supercoiling is an extra twist added on top of DNA’s normal double helix. This higher-order coiling helps pack a very long DNA molecule into a chromosome, allowing it to fit inside the nucleus.
Chromatin vs. chromosome
Chromatin is DNA plus its associated proteins (histones and other chromosomal proteins). When this material becomes highly condensed and visible under a microscope (such as during mitosis), it’s called a chromosome.
During interphase, eukaryotic chromosomes contain two regions that differ in packing density and typical activity:
- The densely packed heterochromatin - often found at centromeres and telomeres - generally contains inactive genes.
- The more open euchromatin contains actively transcribed genes. Here, DNA is wrapped around nucleosomes but is not as tightly compacted.
Control of gene expression in prokaryotes
- Prokaryotic gene expression is regulated through both negative and positive mechanisms:
- Negative control, exerted by repressors and co-repressors, reduces transcription.
- Positive control, driven by inducers and enhancers, increases transcription.
This regulation allows prokaryotes to respond quickly to environmental changes by adjusting gene expression precisely.
Transcription factors
- Transcription factors in prokaryotes bind regulatory DNA elements such as enhancers and silencers, typically located near the core promoter within an extended promoter region.
- When activator proteins bind enhancers, they increase transcription. When repressor proteins bind silencers, they decrease transcription.
Operon concept and Jacob-Monod model
- A common prokaryotic regulatory strategy uses operons, which are groups of genes controlled by a single promoter.
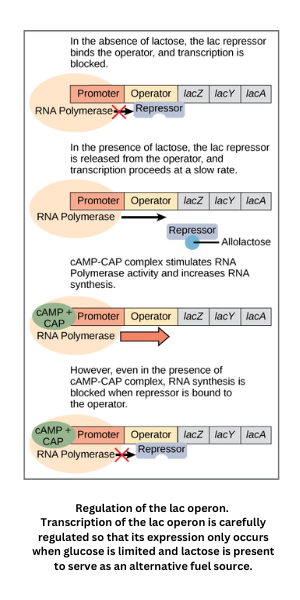
- Within an operon, the operator is the binding site for regulatory proteins. For example, in the lac operon (the Jacob-Monod model), a repressor binds the operator near the promoter and blocks transcription when the appropriate inducer is absent.
Additional regulation involves co-repressors and co-inducers:
- A co-repressor binds its target and converts an otherwise inactive transcription factor into an active repressor.
- A co-inducer can either activate an inducer or inactivate a repressor, further fine-tuning transcription.
Gene repression in bacteria
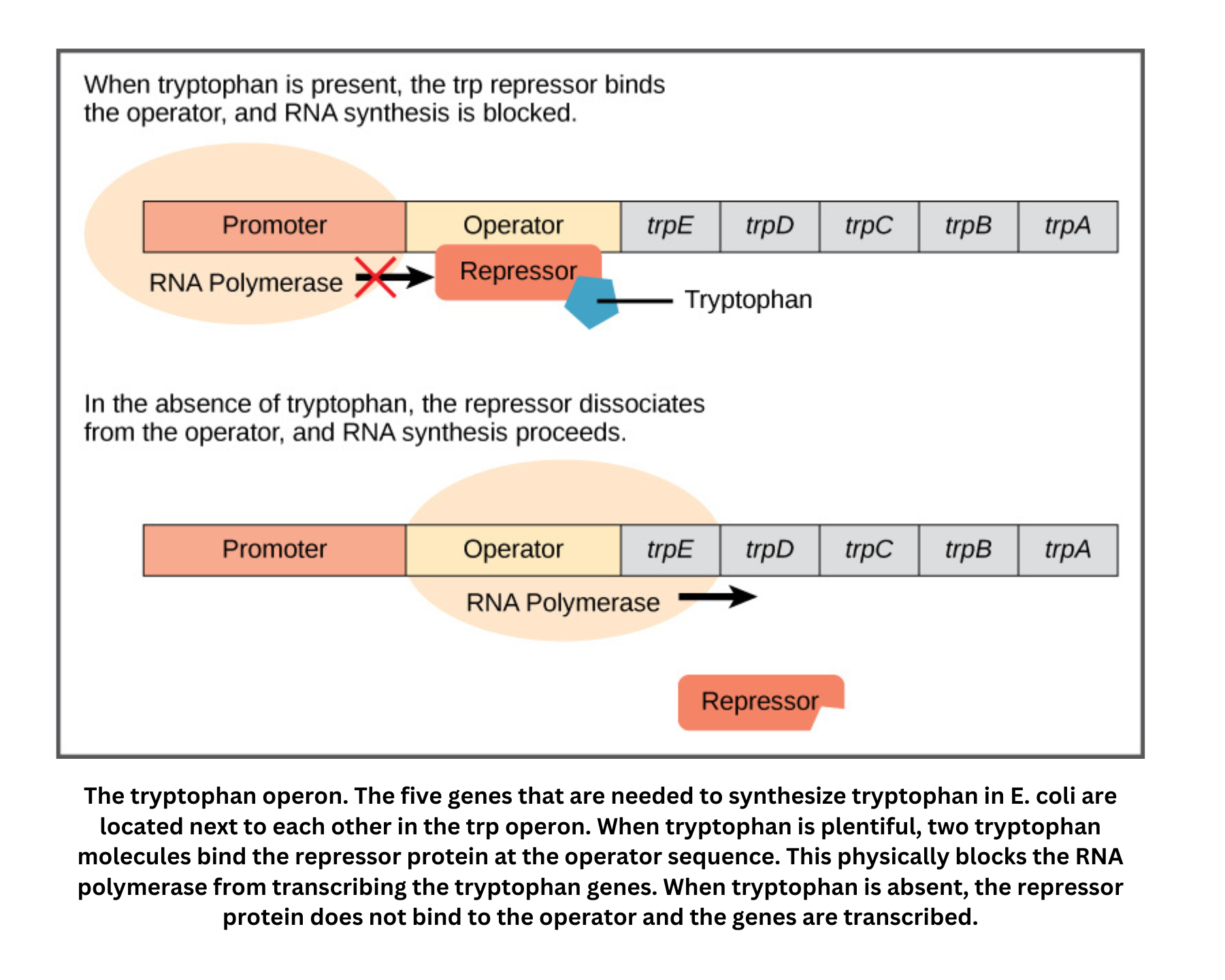
- Transcription attenuation is another regulatory strategy. In the trp operon, a particular stem-loop structure in the mRNA forms when tryptophan levels are high, causing premature termination of transcription. When tryptophan levels are low, transcription continues to produce a full-length transcript.
Positive control in bacteria
- Bacteriophages can redirect the host’s transcription machinery by producing specialized α factors. These factors are made at different stages of infection, ensuring that early, middle, and late phage genes are transcribed in the correct order.
Control of gene expression in eukaryotes
In eukaryotic cells, gene expression is regulated mainly at the transcriptional and post-transcriptional levels. Unlike prokaryotes - which often use operons and attenuation - eukaryotes rely on mechanisms such as transcription factors, DNA looping, chromatin modifications, and mRNA processing to control when and how strongly genes are expressed.
Transcription regulation
Transcription factors are DNA-binding proteins that change transcription rates by binding to enhancers (activating sequences) or silencers (repressive sequences). These regulatory regions may be far from the promoter (upstream or downstream), so the DNA often must loop to bring the bound transcription factor into contact with the promoter complex. Intermediary proteins (co-activators or co-repressors) help stabilize these interactions and influence RNA polymerase activity.
DNA-binding domains in transcription factors typically interact with the major and minor grooves of the DNA double helix. Common binding motifs include the helix-turn-helix, zinc finger, and basic-region leucine zipper.
Gene amplification and duplication
Eukaryotes can also change expression levels through gene amplification or gene duplication, which increases the number of copies of a gene. This may happen through duplication of chromosomal segments or extra replication of entire chromosomes. Amplified genes such as MYC or RAS in cancer can drive excessive protein production and affect disease outcomes. Some duplications, such as Her2 in breast cancer cells, are clinically important because they can be targeted by specific therapies (e.g., Herceptin). In other cases, duplicated genes have little functional effect beyond increasing genome size.
Post-transcriptional controls
After transcription, several post-transcriptional processes influence how much protein is produced from an mRNA:
- Splicing: Eukaryotic pre-mRNA contains introns (non-coding segments) and exons (coding segments). Introns are removed and exons are joined to form mature mRNA. Alternative splicing allows one gene to produce multiple mRNA variants (and therefore different proteins).
- 5′ cap and 3′ poly-A tail: A 5′ cap (modified guanine) and a poly-A tail at the 3′ end protect mRNA from degradation and promote export from the nucleus.
- RNA modifications: Some RNAs, especially tRNA and rRNA, contain modified nucleotides that support proper structure and function.
- RNA degradation: Cells also regulate how quickly specific mRNAs are broken down. Changing transcript stability is another way to fine-tune protein production over time.
Cancer as a failure of normal cellular controls
In cancer, cells ignore the regulatory signals that normally limit growth. Normal cells typically stop dividing or initiate apoptosis (programmed cell death) when they detect substantial DNA damage. Cancer cells continue dividing under conditions that would usually halt proliferation. They can also promote angiogenesis (new blood vessel growth to supply nutrients), become “immortal” by bypassing normal division limits, and spread through metastasis, forming secondary tumors in other tissues.
Oncogenes and tumor suppressors
- Oncogenes are genes that, when activated or mutated, drive continuous cell division. Their normal versions, proto-oncogenes, usually help regulate growth and division. When converted into oncogenes (for example, src), they can trigger uncontrolled proliferation.
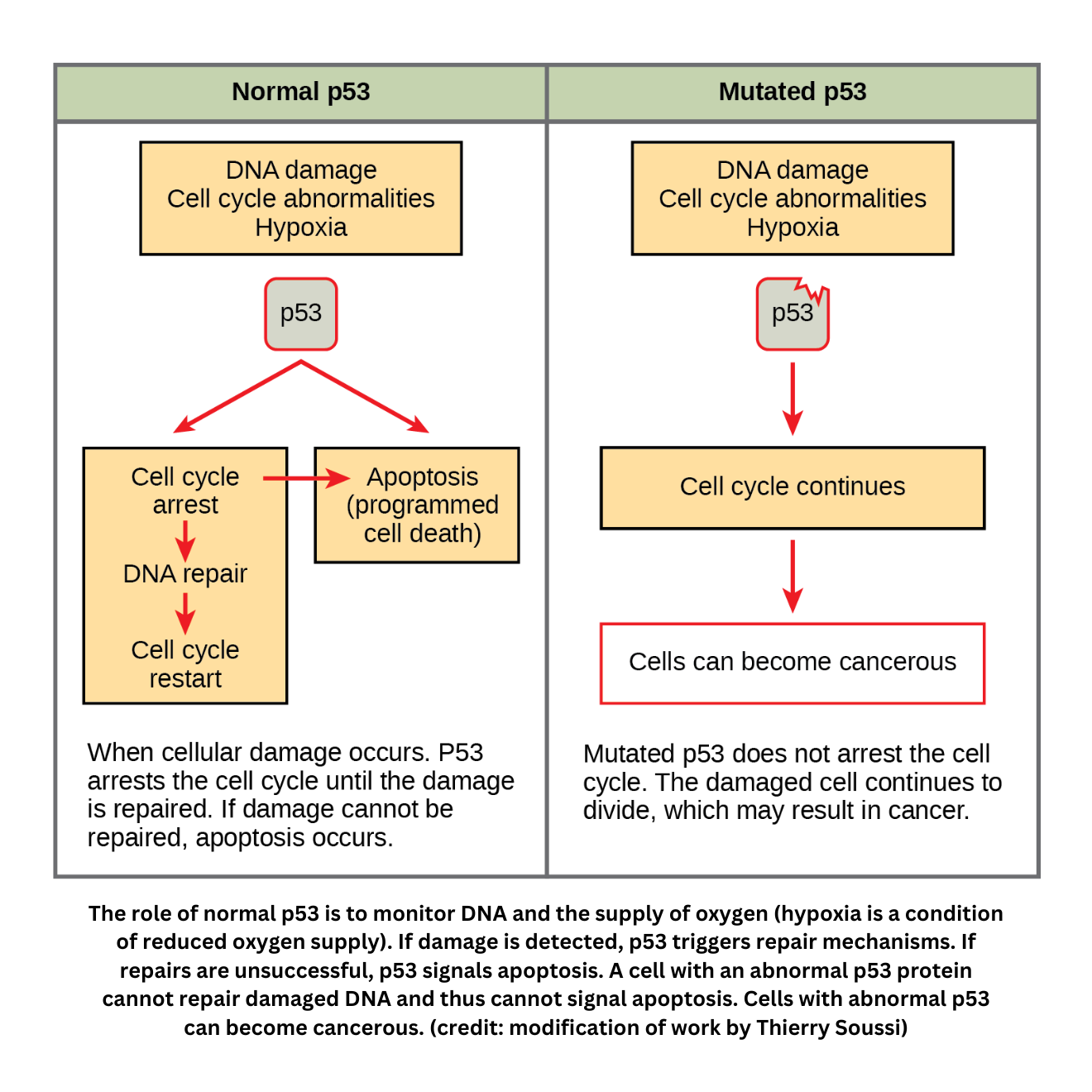
- Tumor suppressor genes counterbalance growth signals by limiting or monitoring cell division. If these genes (such as p53) lose function, abnormal proliferation becomes more likely.
Regulation of chromatin structure
Chromatin is DNA wrapped around histone proteins. How tightly chromatin is packed affects gene expression:
Euchromatin: Loosely packed, which supports active transcription.
Heterochromatin: Tightly packed, which reduces access to DNA.
Cells modulate chromatin by modifying DNA or histones:
- DNA methylation: In eukaryotes, CpG islands (regions rich in cytosine and guanine) near promoters are common methylation targets for DNA methyltransferase (DNMT). Adding methyl groups (for example, forming 5-methylcytosine) typically silences the associated genes.
- Histone modifications: Histone methylation and acetylation can shift chromatin between active (euchromatin) and inactive (heterochromatin) states.
Non-coding RNAs
Non-coding RNAs include all RNA types other than mRNA. They have structural and regulatory roles:
- rRNA (ribosomal RNA): Forms core structural and functional components of ribosomes.
- tRNA (transfer RNA): Delivers amino acids during translation.
- snRNA (small nuclear RNA): Functions in RNA splicing (as part of spliceosomes).
- snoRNA (small nucleolar RNA): Guides chemical modifications of RNA.
- miRNA (microRNA): Participates in RNA silencing by inhibiting translation or promoting mRNA degradation.
Modifications such as 2′-O methylation and conversion of uracil to pseudouracil can stabilize these RNAs, support their structure, and improve their function.