Transcription and translation
Transcription
tRNA and rRNA: composition and roles in translation:
- Although transfer RNA (tRNA) and ribosomal RNA (rRNA) are produced by transcription, they don’t serve as the direct template for translation (unlike mRNA). Instead, tRNA delivers the correct amino acid to the ribosome during protein synthesis, and rRNA forms a major part of the ribosome and helps catalyze peptide bond formation.
tRNA structure and function:
- Nucleotide composition: tRNA contains many chemically modified nucleotides that improve its stability and function.
- Cloverleaf configuration: tRNA typically folds into a cloverleaf shape with an anticodon loop (which base-pairs with the mRNA codon) and an amino acid attachment site at the 3′ end. The amino acid attaches to the tRNA’s 3′-OH group through an ester linkage.
rRNA structure and function:
- Nucleotide composition: rRNA also contains modified nucleotides that support proper folding and catalytic activity.
- Catalytic role: The rRNA in the large ribosomal subunit contains the active site that catalyzes peptide bond formation. Notably, rRNA can catalyze this reaction even without ribosomal proteins, which is why rRNA is considered a ribozyme.
Transcription entails synthesizing RNA from a DNA template and requires:
- RNA polymerase and promoters: RNA polymerase binds a promoter region (e.g., a TATA box) in double-stranded DNA. The DNA is initially closed, then unwinds to form an open complex.
- Chain initiation: No primer is needed (unlike DNA replication). Once RNA polymerase binds the promoter, it begins adding ribonucleotides complementary to the DNA template strand.
- Chain elongation: Nucleoside triphosphates (ATP, UTP, GTP, and CTP) are added in the 5′ 3′ direction, extending the growing RNA strand.
- Chain termination:
- Intrinsic termination: A particular sequence causes the RNA to form a stem-loop structure, which leads to detachment from the DNA template.
- Rho-dependent termination: The ρ factor (rho protein) moves along the nascent RNA, catches RNA polymerase, and dislodges it from the DNA.
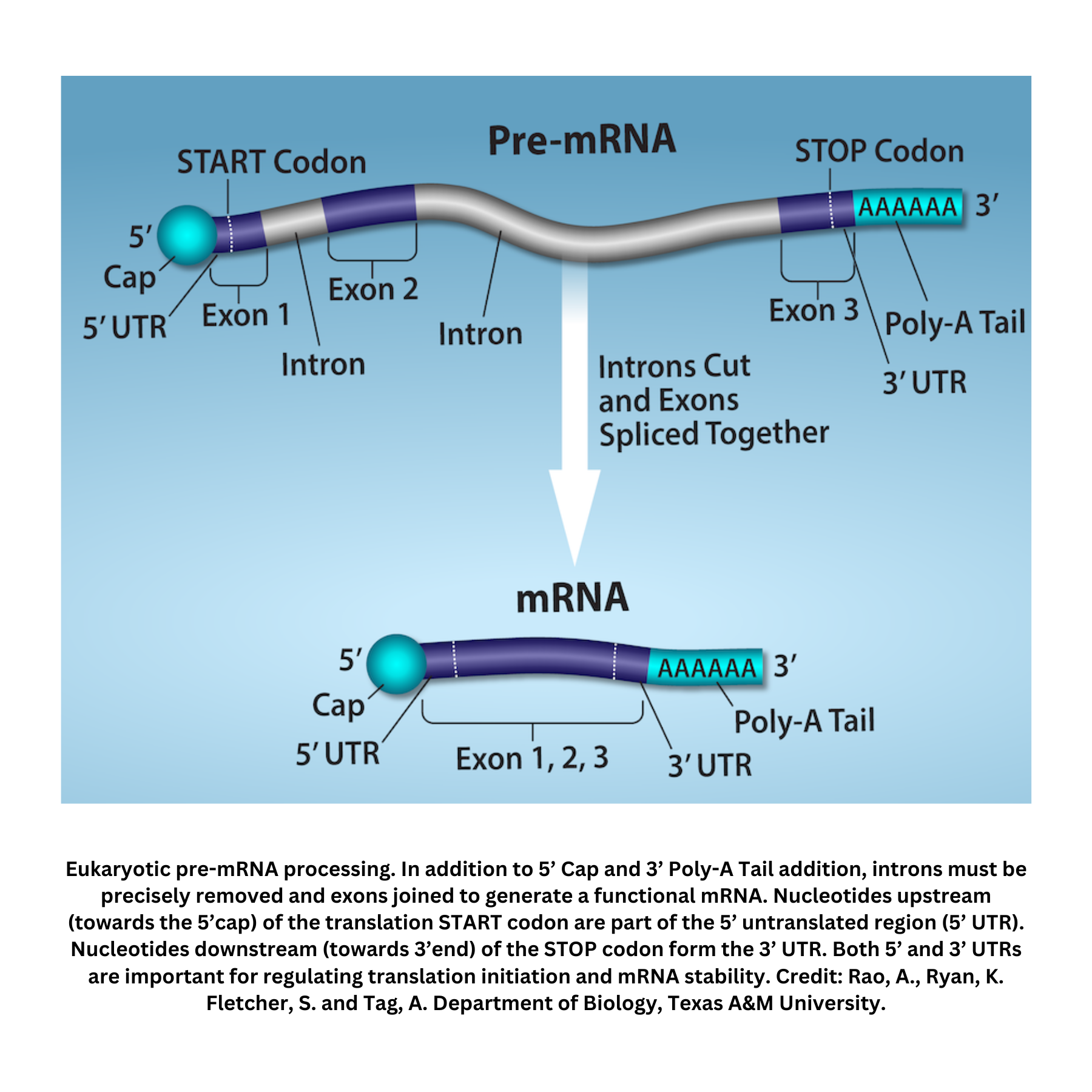
mRNA processing in eukaryotes:
In eukaryotic cells, mRNA undergoes post-transcriptional modifications that stabilize it and prepare it for export to the cytoplasm:
- 5′ cap: A modified guanine nucleotide attached in an unusual linkage at the mRNA’s 5′ end. This helps protect the mRNA from exonuclease degradation and promotes ribosomal binding.
- 3′ poly-A tail: A stretch of adenine residues added to the 3′ end. This helps prevent rapid breakdown and supports nuclear export.
- Splicing: Non-coding regions (introns) are removed, and coding regions (exons) are joined together. Alternative splicing allows a single gene to produce multiple mRNA isoforms.
Prokaryotic mRNA generally lacks the 5′ cap and poly-A tail, and in some bacteria, transcription and translation can occur at the same time.
Ribozymes, spliceosomes, and RNA-associated complexes:
- Ribozymes: RNA molecules that act as enzymes. A key example is the rRNA in the large ribosomal subunit.
- Spliceosomes: Large RNA-protein complexes that carry out mRNA splicing. Their RNA components can contribute ribozyme-like activity.
- snRNPs (small nuclear ribonucleoproteins): RNA-protein subunits that assemble into spliceosomes.
- snRNAs (small nuclear RNAs): The RNA components of snRNPs, which help recognize splice sites and catalyze intron removal.
Translation
Roles of mRNA, tRNA, and rRNA
- Messenger RNA (mRNA) carries genetic information from DNA. Each three-nucleotide codon specifies an amino acid in the growing polypeptide.
- Transfer RNA (tRNA) contains an anticodon that pairs with the matching codon on mRNA and carries an amino acid attached at the 3′ end by an ester linkage. This pairing helps ensure the correct amino acid is added when its codon appears on the mRNA.
- Ribosomal RNA (rRNA) forms the core of the ribosome, the complex that catalyzes peptide bond formation. The large subunit’s rRNA plays the catalytic (ribozyme-like) role in linking amino acids, while the small subunit’s rRNA helps recognize and position the mRNA.
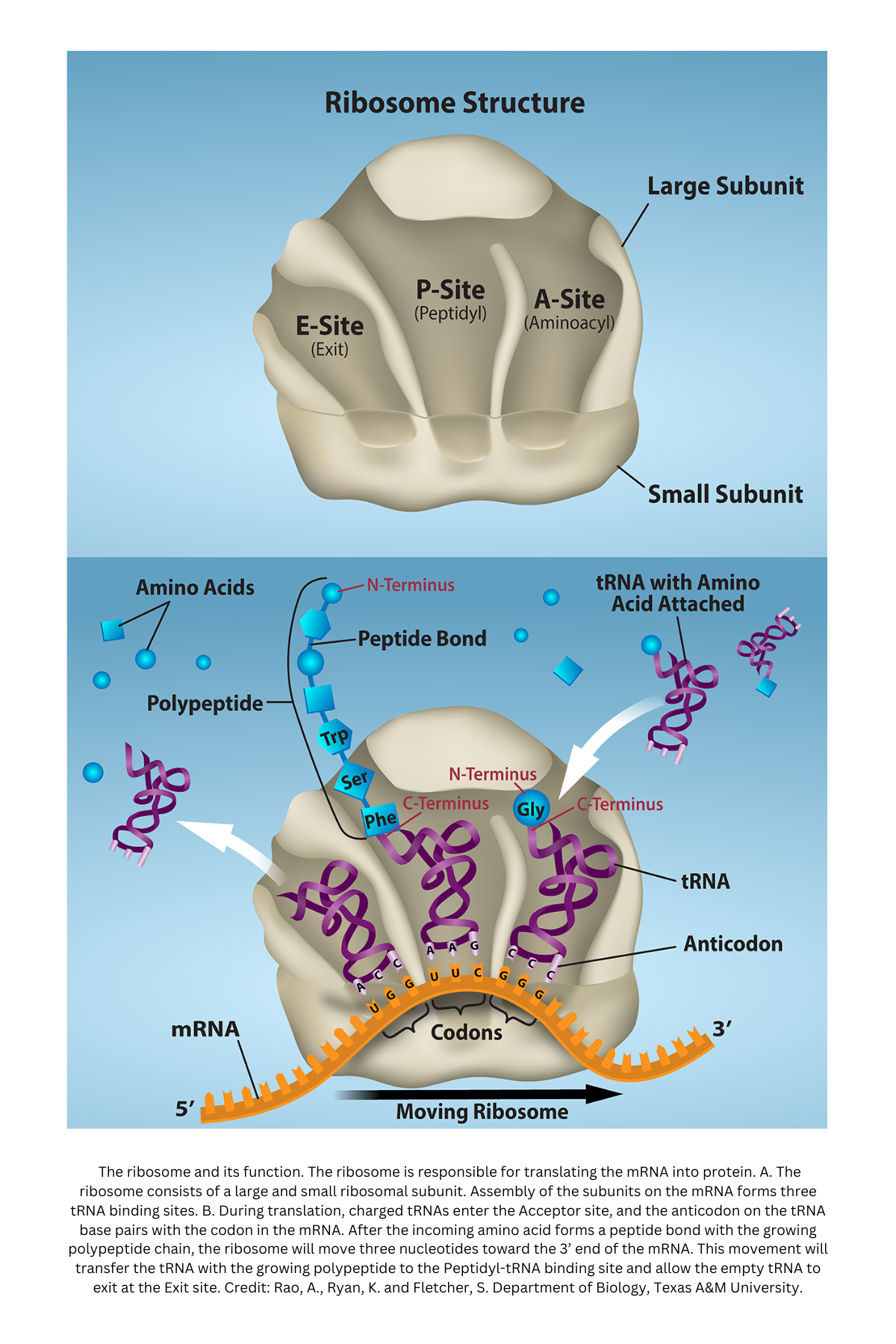
Ribosome structure and function:
A ribosome is the site of translation (protein synthesis). It has two subunits (large and small) that assemble on the mRNA:
- Large subunit: Catalyzes peptide bond formation (peptidyl transfer).
- Small subunit: Binds and positions the mRNA, recognizing sequences such as the Shine-Dalgarno region in prokaryotes or the Kozak sequence in eukaryotes.
When the subunits align, they form a “sandwich” around the mRNA and incoming tRNAs, holding them in place so amino acids can be added in the correct order.
Mechanism of translation: initiation, elongation, and termination:
- Chain initiation:
- In prokaryotes, the small ribosomal subunit binds near the Shine-Dalgarno sequence on the mRNA (in eukaryotes, it targets the Kozak sequence).
- An initiator tRNA (often carrying formyl-methionine, fMet, in prokaryotes) binds the start codon (AUG).
- Initiation factors and GTP help assemble the complete ribosome, positioning the initiator tRNA in the P site to begin translation.
- Chain elongation:
- Aminoacyl-tRNA binding: A charged tRNA (an aminoacyl-tRNA) enters the A site with help from elongation factors and GTP.
- Peptidyl transfer: The large subunit catalyzes peptide bond formation between the growing chain in the P site and the amino acid in the A site. This transfers the growing peptide to the A-site tRNA.
- Translocation: The ribosome moves one codon along the mRNA. The peptidyl-tRNA shifts from the A site to the P site, the empty tRNA moves to the E site and exits, and the A site opens for the next aminoacyl-tRNA.
- Chain termination:
- When a stop codon (UAA, UAG, or UGA) enters the A site, release factors (bound to GTP) bind instead of a tRNA.
- The completed polypeptide is released from the final tRNA in the P site, and the ribosome dissociates.
Amino acid activation:
- Before translation begins, aminoacyl-tRNA synthetases attach the correct amino acid to its corresponding tRNA. This reaction uses ATP and ensures each tRNA is charged with the appropriate amino acid.
Post-translational modification of proteins
After translation, proteins often undergo additional modifications, including:
- Glycosylation: Adding carbohydrate groups for targeting or structural roles (e.g., in proteoglycans).
- Lipid addition: Adding fatty groups for membrane anchoring.
- Acetylation: A modification that can direct subcellular localization.
- Disulfide bond formation: Linking parts of a protein (or separate subunits) to stabilize tertiary or quaternary structure.
- Phosphorylation: Activating or deactivating enzymes.
- Ubiquitination: Marking proteins for degradation.