Nucleic acid structure, replication and repair
DNA and RNA structure
- DNA stands for deoxyribonucleic acid
- Deoxy = “without oxygen” because DNA lacks an oxygen at the 2’ ring carbon position in the ribose sugar (just )
- RNA stands for ribonucleic acid and has at the 2’ ring carbon position in the ribose sugar
- Ribo = ribose (pentose) sugar
- The acidity in DNA and RNA comes from the phosphate group.
- Nucleic acids are macromolecules essential for storing and transmitting genetic information.
Nucleotides
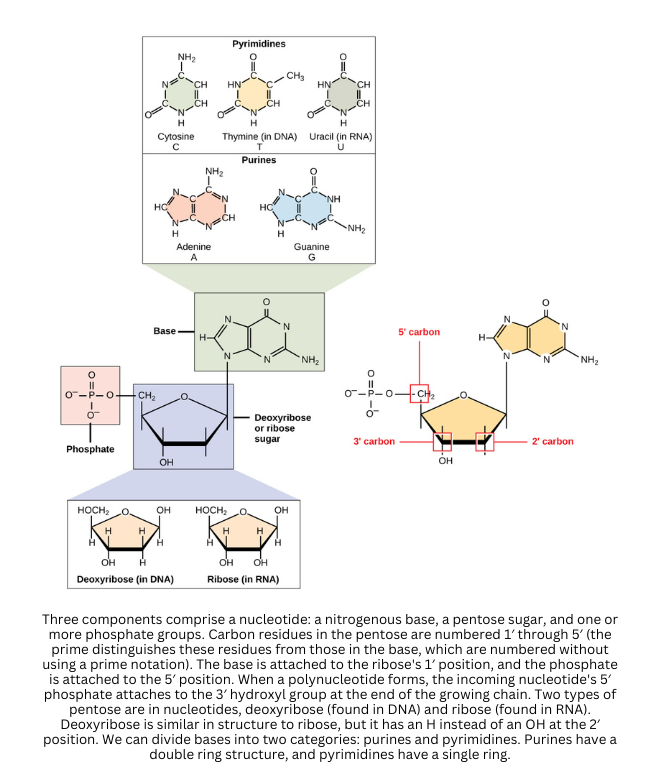
- The fundamental units of nucleic acids are nucleotides. Each nucleotide contains a five-carbon sugar, a phosphate group, and a nitrogenous base. In contrast, nucleosides contain only the sugar and base, so they lack the phosphate.
When multiple nucleotides link together through phosphodiester bonds (between the sugar of one nucleotide and the phosphate of the next), they form the sugar-phosphate backbone characteristic of DNA and RNA.
Attached to this backbone, the nitrogenous bases project inward. These bases fall into two main groups:
- Purines (adenine, guanine) have a double-ring structure
- Pyrimidines (cytosine, thymine, uracil) have a single ring
In DNA - or deoxyribonucleic acid - thymine is used instead of uracil. In most cells, DNA is typically double-stranded. According to the Watson-Crick model (based heavily on Rosalind Franklin’s images), DNA forms a double helix, where two complementary strands wrap around a common axis.
DNA’s primary role is the transmission of genetic information across generations. The sequence of bases encodes instructions that direct cellular processes. The two strands run in opposite directions (they’re antiparallel), which allows each strand to serve as a template during replication.
Under conditions such as high temperature or extreme pH, DNA can undergo denaturation. In denaturation, hydrogen bonds break and the two strands unwind. If optimal conditions return, reannealing can occur: complementary bases realign, rebond, and the helix re-forms.
Similarly, hybridization uses base complementarity by allowing single-stranded DNA from different sources - or DNA with RNA - to base-pair when matching sequences are present.
DNA replication
DNA replication is the coordinated process a cell uses to duplicate its genetic material. Starting from one original double-stranded DNA molecule, replication produces two identical copies of double-stranded DNA.
Replication begins as the helix is opened:
- DNA gyrase helps uncoil DNA in front of the replication fork.
- Helicase unwinds the double helix at the replication fork.
- Unwinding occurs at specific sites called origins of replication. In eukaryotes, there are many origins along each chromosome, which speeds up replication.
- Single-strand binding protein (SSB) binds exposed single-stranded DNA to stabilize it and prevent reannealing.
Once the strands are separated, each original strand serves as a template for a new complementary strand:
- Free nucleotides pair with exposed bases using base-pairing rules (A with T, G with C).
- DNA synthesis always proceeds in the 5′ → 3′ direction.
- Because the strands are antiparallel, the template is read in the 3′ → 5′ direction.
To start synthesis, the cell needs a short starting segment:
- Primase lays down a short RNA primer that base-pairs with the DNA template.
- Later, the RNA primer is replaced with DNA.
- DNA polymerase extends from the primer, adding nucleotides to build the complementary strand.
Both strands are copied at the same time, but they’re made differently because of the 5′ → 3′ rule:
- The strand synthesized in the same direction as fork movement is the leading strand.
- The strand synthesized opposite the fork movement is the lagging strand, made in short segments called Okazaki fragments.
To finish the lagging strand:
- A specialized DNA polymerase removes RNA primers and replaces them with DNA.
- DNA ligase seals the remaining gaps between Okazaki fragments.
Replication proceeds in both directions from each origin, producing two replication forks that move outward and form a replication “bubble.” DNA polymerase also has a proofreading function that corrects most errors during synthesis. In eukaryotes, replication occurs once per cell generation during S phase. Although meiosis involves two rounds of cell division, the replication of DNA itself occurs only once.
DNA polymerase enzymes also catalyze the formation of phosphodiester bonds that link nucleotides into the growing DNA chain.
Semi-conservative nature of DNA replication
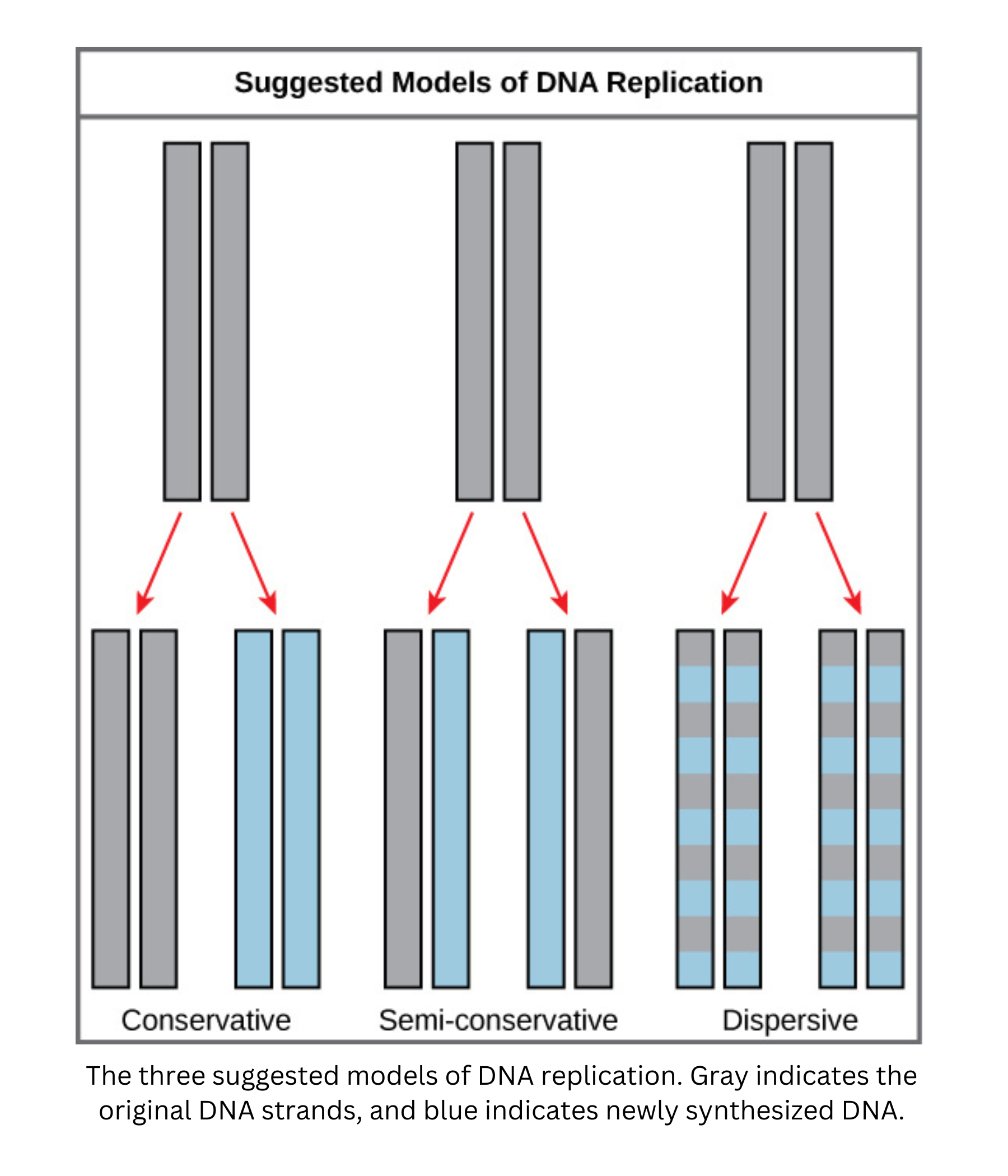
Each new DNA double helix contains one parental strand and one newly synthesized (daughter) strand. For that reason, replication is described as semi-conservative.
Meselson and Stahl designed an experiment to distinguish among models of DNA replication. They grew bacteria in a medium containing a heavy nitrogen isotope (15N), so all bacterial DNA became labeled with heavy nitrogen. They then moved the bacteria to a medium containing only the lighter isotope (14N).
After one round of replication, they extracted the DNA and used density gradient centrifugation. The results did not show two distinct bands (as expected for conservative replication) and did not show a single band that gradually shifts each generation (as expected for dispersive replication). Instead, they observed a single band with intermediate density. This indicated that each DNA molecule contained one heavy (old) strand and one light (new) strand, supporting the semi-conservative model.
After a second round of replication, two bands appeared: one at the intermediate (hybrid) density and a new, lighter band. The lighter band showed that some DNA molecules now had two newly synthesized strands, while the hybrid molecules still contained one old and one new strand.
The replication forks advance in both directions away from each origin, creating replication bubbles.
A special challenge occurs at the ends of linear DNA in eukaryotes, called telomeres. Over repeated rounds of replication, telomeres can shorten because DNA polymerase can’t fully replicate the extreme 3′ end of the lagging strand. Telomerase extends telomeric regions, helping solve the end-replication problem and maintain chromosome integrity across many cell divisions. Over a lifetime, gradual telomere shortening is associated with physiological aging.
Repair of DNA
Repair during replication
- During DNA replication, DNA polymerase reduces errors using a proofreading mechanism called 3′→5′ exonuclease activity. If an incorrect nucleotide is inserted, the enzyme detects the mismatch, reverses direction (“backs up”), excises the incorrect base, and inserts the correct one.
- A different specialized polymerase replaces RNA primers with DNA. It uses 5′→3′ exonuclease activity to remove primers (or short stretches of incorrect nucleotides) and then fills in the region with the correct DNA sequence.
Repair of mutations
- Other repair pathways correct DNA damage and mutations that occur outside replication or alongside it:
- Base-excision repair: Removes a single damaged base (often from deamination or other minor lesions) along with some nearby nucleotides. DNA polymerase fills the gap, and ligase seals it to restore the original sequence.
- Mismatch repair: Enzymes scan for incorrectly paired bases. When a mismatch is found, they excise the region containing the error, and a polymerase resynthesizes the missing section. In organisms such as E. coli, repair enzymes identify the newly synthesized (unmethylated) strand versus the parental (methylated) strand, so only the incorrect strand is cut.
- Nucleotide-excision repair: Fixes bulky lesions (for example, UV-induced thymine dimers) by removing a larger stretch of DNA around the damage. Polymerase replaces the missing segment, and ligase seals it. This pathway handles larger distortions than mismatch or base-excision repair.
- Nick translation: A specialized 5′→3′ exonuclease function coupled to polymerase activity. The enzyme removes defective or RNA-based nucleotides while simultaneously replacing them with correct DNA, as when RNA primers are removed during replication.
- SOS response in some bacteria (notably ): When DNA damage is extensive and overwhelms normal repair, the cell uses a more error-prone polymerase to continue replication across damaged templates. This can introduce mutations, but it prevents replication from stalling completely, which could otherwise be lethal to the cell.