Recombinant DNA and biotechnology
Recombinant DNA and its applications
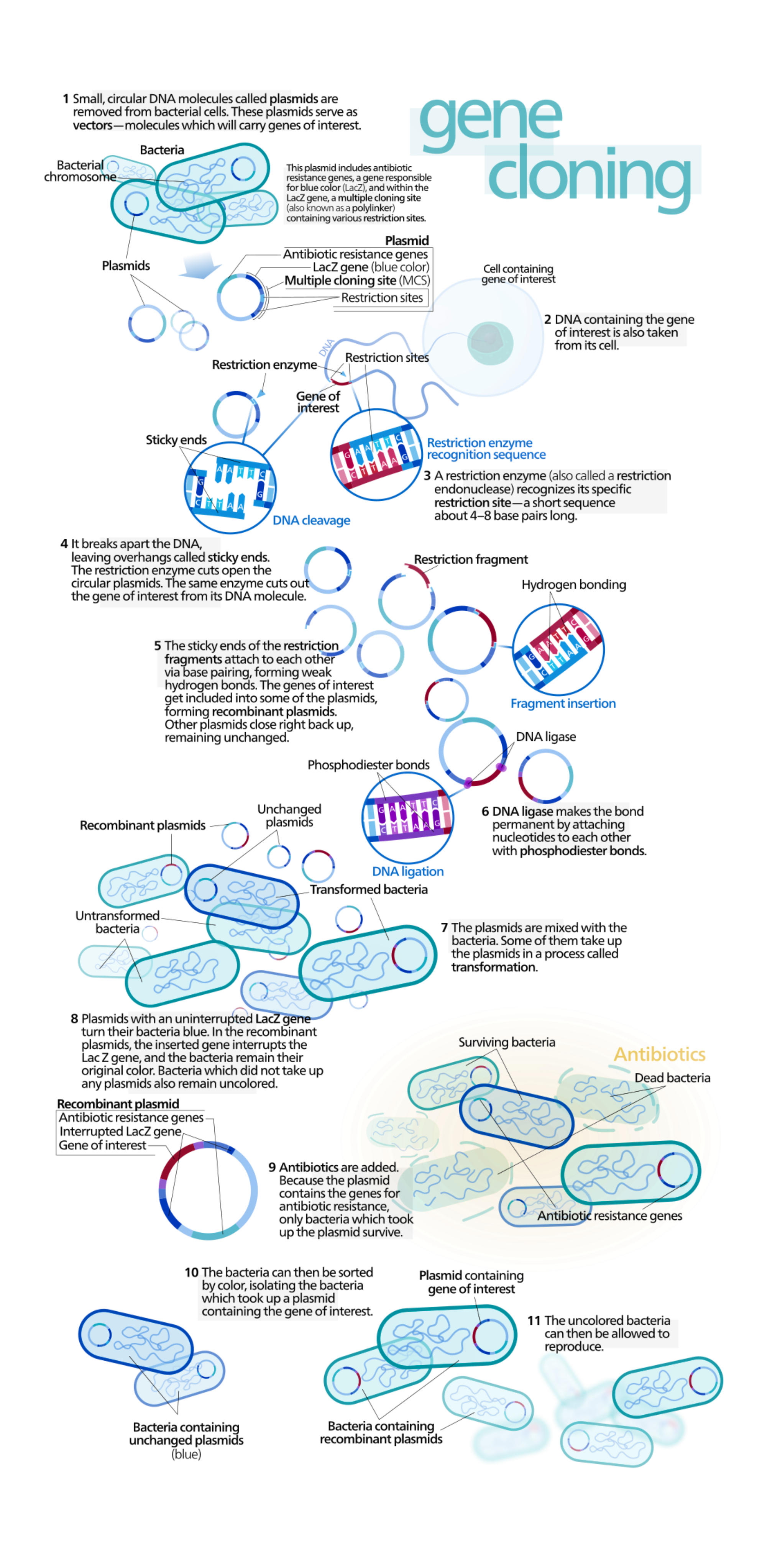
Gene cloning and plasmids
A plasmid is often used for gene cloning because it has three useful features:
- A restriction site that can be cut by a restriction enzyme, creating an opening where the gene of interest can be inserted.
- An origin of replication, which lets the plasmid (and the inserted gene) replicate independently of the host cell’s genome.
- An antibiotic resistance gene, so only bacteria that took up the plasmid survive when grown with that antibiotic.
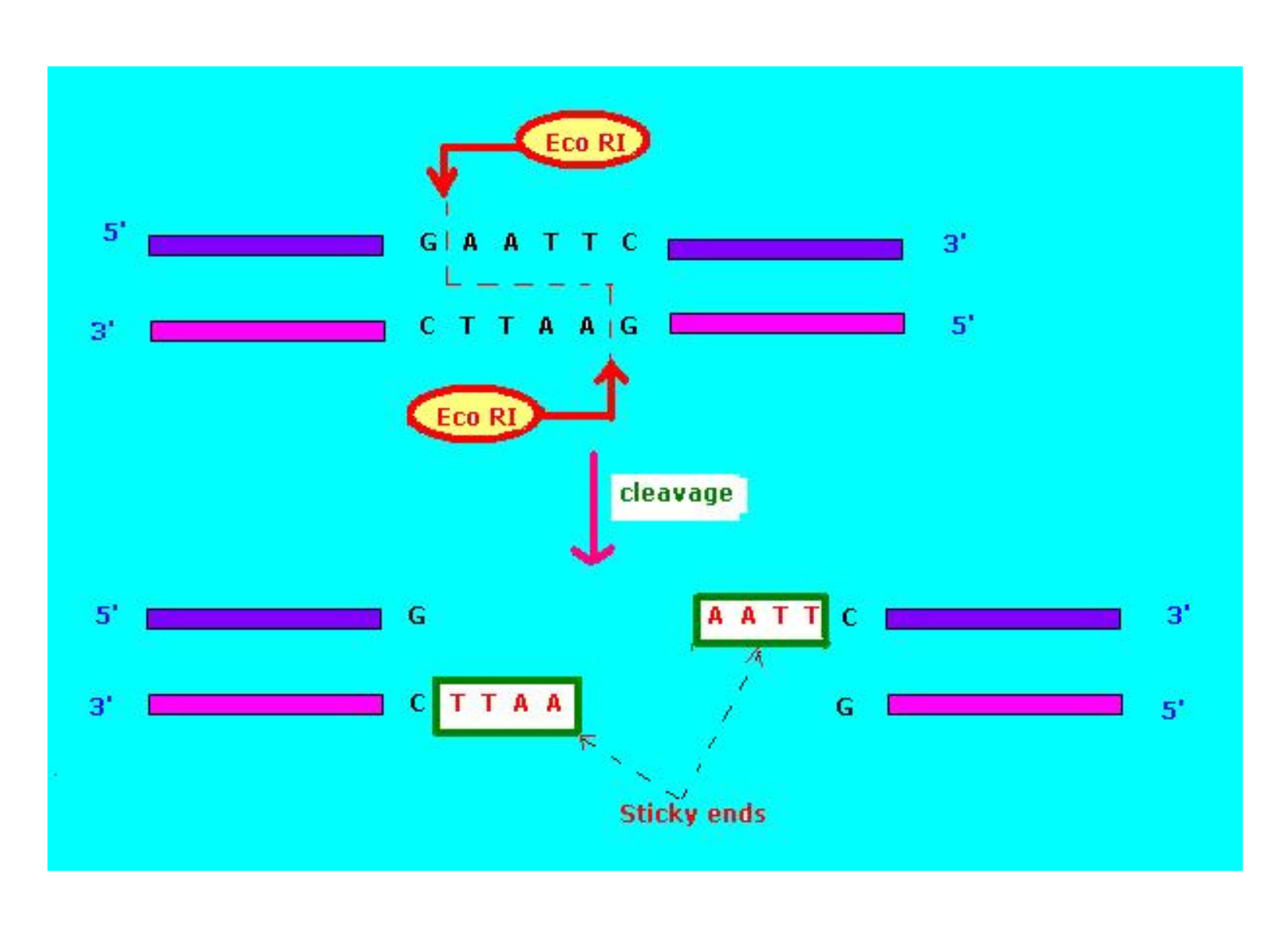
During cloning, restriction enzymes (endonucleases) cut double-stranded DNA at palindromic sequences. Depending on the enzyme, the cut produces either:
- Sticky ends, which have single-stranded overhangs that can base-pair (reanneal) with complementary sequences
- Blunt ends, which have no overhangs and therefore don’t base-pair with each other
After the gene fragment is joined to the plasmid, the recombinant plasmid is introduced into host cells. As the host cells replicate, they also replicate the plasmid, which clones the inserted gene.
DNA libraries
Researchers generate DNA libraries by cloning a large collection of DNA fragments from an organism:
- Genomic library: Created by fragmenting all genomic DNA with restriction enzymes and cloning the pieces into vectors.
- cDNA library: Made by reverse-transcribing the mRNA population into complementary DNA (cDNA), then cloning those cDNA fragments. This library reflects only expressed genes.
Hybridization
- Hybridization (annealing) is the binding of single-stranded DNA molecules to complementary sequences.
- In Southern blot analysis, a DNA probe anneals to a matching target DNA fragment on a gel.
- In gene cloning, sticky ends on a gene fragment can hybridize with complementary sticky ends on a plasmid.
PCR (polymerase chain reaction)
PCR amplifies a specific DNA region exponentially by repeating three steps:
- Denaturation: Heat separates the two DNA strands.
- Annealing: Cooling allows primers to bind to their complementary sequences.
- Elongation: A heat-stable polymerase extends each primer in the 5′→3′ direction.
Because the amount of target DNA doubles each cycle, amplification follows .
Gel electrophoresis and blotting
- Gel electrophoresis separates DNA fragments by size as they move through a gel matrix.
- Southern blot uses a labeled DNA probe to detect a specific DNA fragment on the gel.
- Other blots include Northern (RNA target, DNA or RNA probe) and Western (protein target, antibody probe).
DNA sequencing
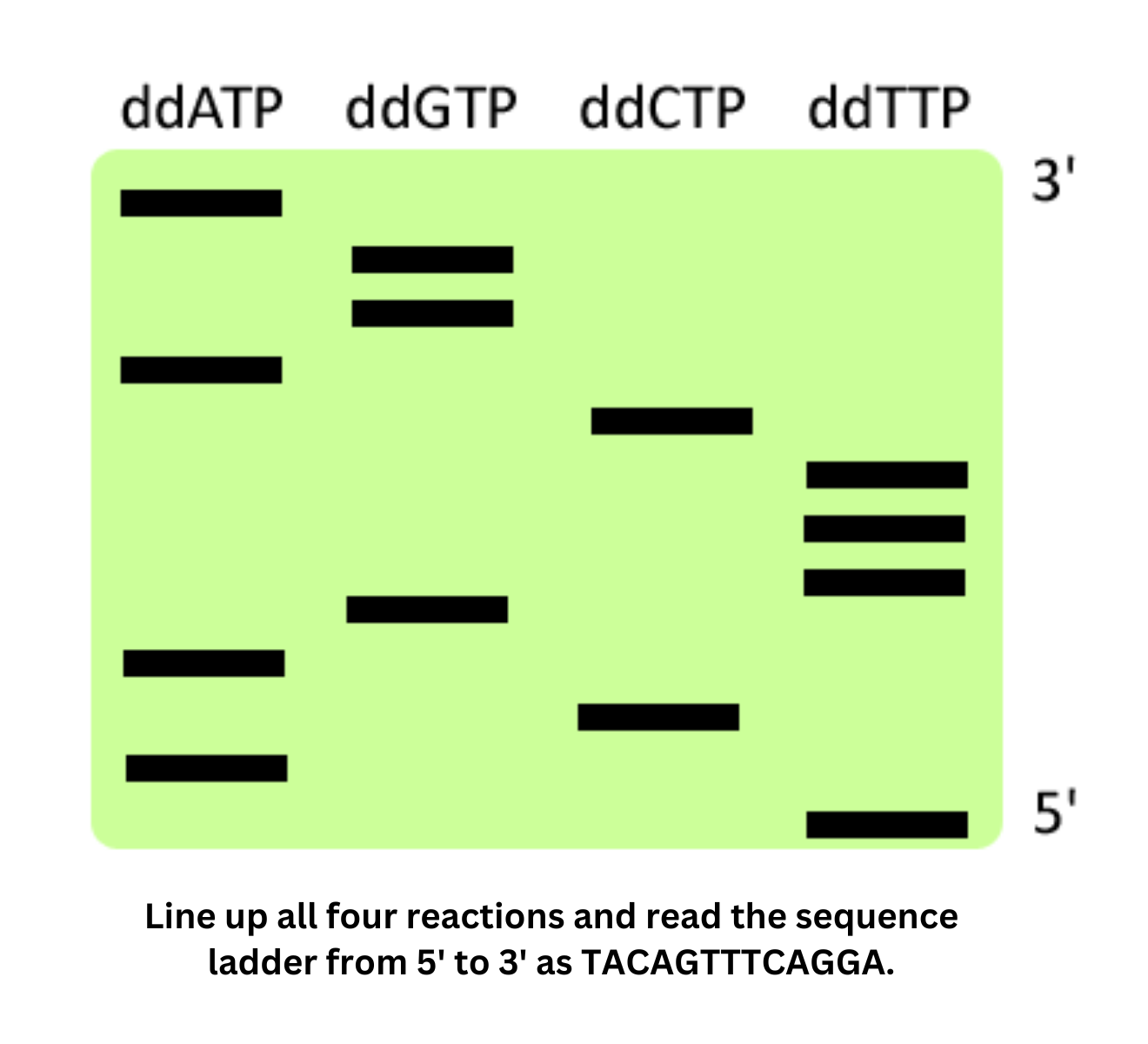
- Sanger sequencing uses dideoxy-NTPs to terminate DNA synthesis at specific bases, and the resulting fragments can be read by capillary electrophoresis. It’s economical for short sequences but limited in scope.
- Next-generation sequencing runs many sequencing reactions in parallel. It’s useful for whole genomes or detecting low-level variants, but it’s more expensive.
Gene expression analysis
- mRNA levels are commonly used as a readout of gene expression.
- Microarray (gene expression profiling) shows which genes are overexpressed or underexpressed across the genome (for example, in cancer).
- qRT-PCR (quantitative reverse transcription PCR) measures expression of a specific gene, such as residual BCR-ABL transcripts in leukemia.
Stem cells
- Stem cells can differentiate into many cell types.
- For example, hematopoietic stem cells can produce multiple types of blood cells.
- Clinically, a bone marrow transplant uses these stem cells to treat leukemia.
Practical applications of DNA technology
- Medical: PCR plus sequencing can diagnose mutations, predict drug responses, or monitor diseases.
- Gene therapy uses viruses to insert functional genes into human cells.
- Pharmaceuticals: Recombinant proteins or enzymes can treat deficiencies.
- Forensics: DNA fingerprinting can be used for paternity testing or criminal identification.
- Environmental: Genetically engineered bacteria can be used to degrade oil spills or plastics.
- Agriculture: GMO crops can be engineered for improved yields or pest resistance.
Safety and ethics
- Gene therapy can inadvertently introduce mutations that cause cancer.
- Genetically modified foods may reduce the diversity of edible plant species and increase susceptibility in the food supply, and their possible negative long term effects on human health have not been absolutely ruled out.
- Full genome sequencing raises privacy and discrimination concerns regarding genetic data.
Determining gene function
Understanding how a particular gene contributes to biological processes often involves combining comparative genomics, protein domain analysis, interaction studies, and expression profiling. Each approach provides a different kind of evidence, and together they help clarify a gene’s role in the cell or organism.
- Evolutionary comparison
- Cross-species conservation: Researchers ask whether a gene is conserved across diverse organisms. Genes shared across multiple species, especially at the protein level, often carry out fundamental functions. Sequence similarities can also highlight regions that are critical for activity or stability.
- Phylogenetic analysis: By reconstructing evolutionary relationships, scientists can estimate when a gene (or its variants) first appeared, which can suggest its earliest function and why it was retained.
- Protein domains
- Domain architecture: Many proteins contain recognizable structural segments (for example, kinase domains or DNA-binding motifs). Identifying these domains helps predict how the protein interacts with other molecules and whether it’s likely involved in signaling, catalysis, or nucleic acid binding.
- Functional predictions: If a newly discovered protein shares a domain with a well-studied protein family, it may perform similar tasks - even in a different organism.
- Protein interactions
- Interaction networks: Identifying binding partners can place a protein in its functional context. Techniques such as co-immunoprecipitation, yeast two-hybrid, or mass spectrometry-based proteomics can reveal interaction partners that co-regulate pathways or assemble into protein complexes.
- Pathway assignment: If a protein interacts with several known components of a metabolic or signaling cascade, that pattern suggests it participates in that pathway.
- Cellular expression
- Spatial and temporal patterns: Measuring when and where mRNA or protein is present helps show where and when a gene is active. Techniques like in situ hybridization, immunofluorescence, and reporter assays can reveal tissue-specific or developmental-stage-specific expression.
- Perturbations and phenotypes: If changing expression - through knockdown, knockout, or overexpression - produces a consistent change in cellular or organismal traits, that phenotype can point to the gene’s normal role.