Enzyme structure, function and activity
Enzyme structure and function
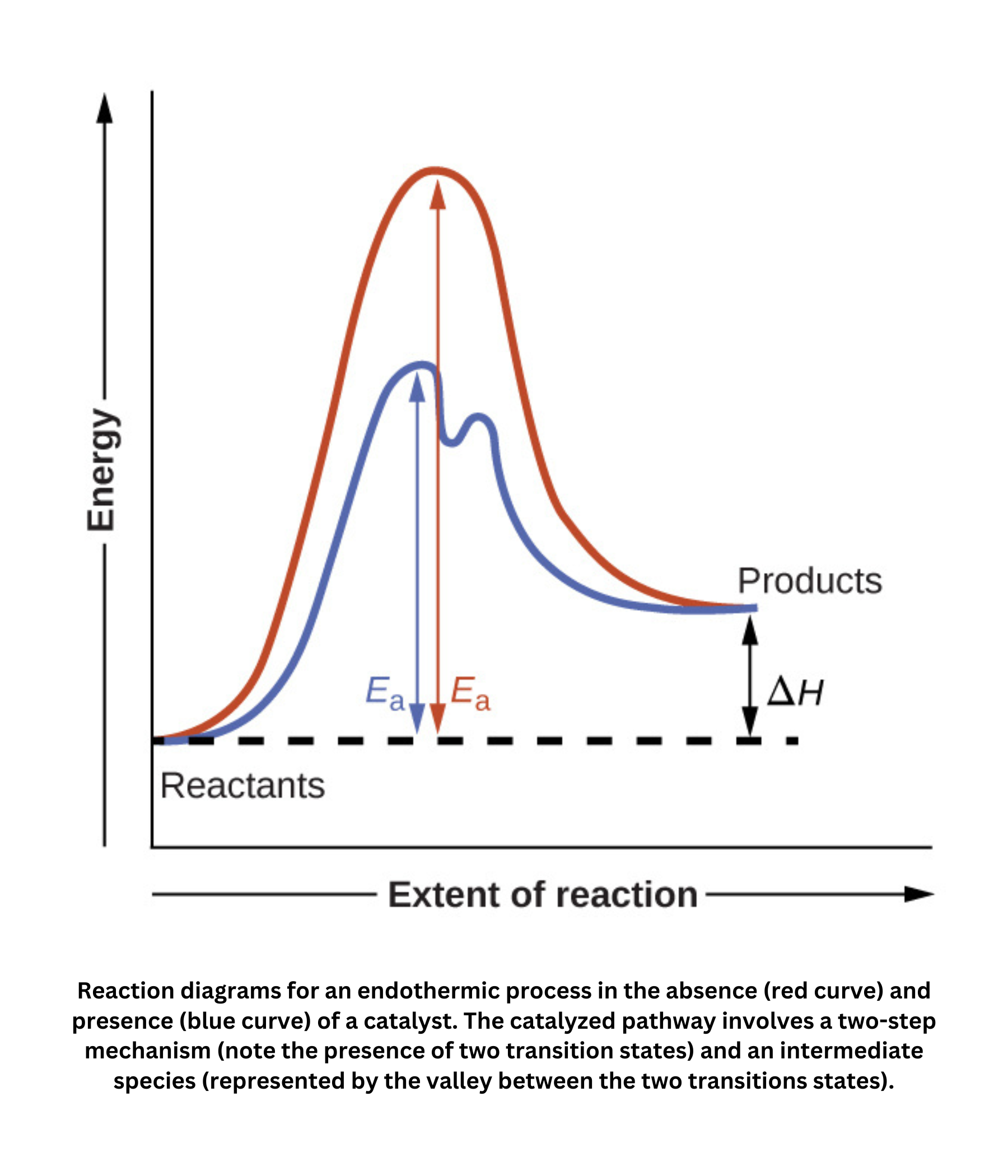
Enzymes are specialized proteins that act as biological catalysts, greatly accelerating chemical reactions in living organisms by lowering the activation energy. This allows essential processes to occur at temperatures and pH levels compatible with cellular life, including:
- Metabolism
- DNA synthesis
- RNA synthesis
- Protein synthesis
- Digestion
Enzymes DO:
- …increase the rate constant, , for the equation rate =
- …speed up both forward and reverse reactions
- …affect a reaction’s kinetics, but not the thermodynamics.
Enzymes DO NOT:
- …alter the of a reaction (because lowers activation energy for forward/reverse)
- …change , the net change in free energy
Composition
- Enzymes are composed of either protein (as in the vast majority of enzymes in the human body) or RNA, with the most important human RNA-based enzyme being the ribosome.
Classification by reaction type
Enzymes are often grouped by the type of reaction they catalyze. Examples include:
- Oxidoreductases, which handle redox transformations
- Transferases, which relocate functional groups
- Hydrolases, which cleave bonds using water
- Lyases, which break bonds without hydrolysis or oxidation
- Isomerases, which rearrange molecular structures
- Ligases, which form new bonds, typically driven by ATP.
Substrates and specificity
Each enzyme selectively binds one or more substrates - the molecules undergoing reaction - based on precise three-dimensional complementarity. This specificity comes from the structural match between the substrate and the enzyme’s active site, which helps align reactants and promotes efficient chemical conversion.
Enzymes typically show high specificity, recognizing subtle differences in substrate shape or functional groups, and they can be specific enough to distinguish stereoisomers.
Mechanism of catalysis and role of cofactors
Enzymes use several strategies to facilitate reactions:
- They may bring substrates into close proximity, stabilize charged intermediates, or distort bonds to better resemble the reaction’s transition state.
- Some enzymes require cofactors or coenzymes - non-protein components that assist in catalysis.
- Cofactors can be metal ions (e.g., zinc, iron), while coenzymes are often carbon-based molecules such as vitamins or their derivatives.
- The water-soluble vitamins (e.g., B-complex vitamins) frequently act as coenzyme precursors, forming essential components like NAD⁺ or FAD.
- These auxiliary components can shuttle electrons or chemical groups, enabling reactions that amino acid side chains cannot perform on their own.
Effects of local conditions
Enzyme activity is strongly influenced by environmental factors such as pH, temperature, and the concentrations of substrates or products. Each enzyme has an optimal pH range and temperature window.
When conditions deviate from this range, hydrogen bonding can be impaired and protein folding can be disrupted, decreasing activity or causing denaturation.
In cells, compartmentalization and regulatory molecules help control these conditions so enzymes function at the right pace and in the right location.
Control of enzyme activity
Enzyme activity is regulated by several mechanisms that fine-tune how rapidly a catalyzed reaction proceeds, allowing cells to adapt to changing conditions.
A key area of study is kinetics, which examines how reaction rates change with substrate concentration, enzyme concentration, and other factors. A classic model used to describe these relationships is the Michaelis-Menten framework, which shows how an enzyme’s velocity increases with substrate concentration until it reaches a maximum rate.
Some enzymes exhibit cooperativity, meaning substrate binding at one subunit or site changes the affinity of other subunits or sites, producing a more dynamic response curve.
Feedback regulation
Beyond intrinsic kinetic behavior, feedback regulation modulates enzyme function at the pathway level. Here, the end product of a pathway binds to upstream enzymes - often the first committed step - to slow or halt its own production. This negative feedback loop helps maintain metabolic balance.
Inhibition
Another layer of control comes from inhibition, in which molecules bind an enzyme and reduce or prevent its activity:
- Competitive inhibitors compete directly with the substrate for the active site, raising the effective substrate concentration needed to achieve a given reaction rate.
- Non-competitive inhibitors bind elsewhere on the enzyme, affecting activity regardless of how much substrate is present.
- Mixed inhibitors can bind either the free enzyme or the enzyme-substrate complex, exerting more nuanced effects.
- Uncompetitive inhibitors predominantly associate with the enzyme-substrate complex, shifting the reaction profile in a unique way.
Regulatory enzymes
Some regulatory enzymes have specialized structural features or undergo modifications that change their catalytic behavior.
Allosteric enzymes contain sites separate from the active site, where binding of an effector molecule changes the enzyme’s shape and function, either activating or inhibiting it.
Others are covalently-modified by processes such as phosphorylation, methylation, or acetylation, which can quickly switch an enzyme on or off.
A zymogen is an inactive precursor form of an enzyme that requires a specific biochemical change - such as cleavage of a peptide fragment - to become active.